| 0. Reference |
0° |
255,128,128 |
965,949,784 |
Composite of all amino acids |
 |
| 1. Histidine |
329° |
255,128,193 |
33,044,533 |
Group IV: Basic amino acids |
 |
| 2. Glutamic acid |
16° |
255,162,128 |
34,954,870 |
Group III: Acidic amino acids |
 |
| 3. Aspartic acid |
31° |
255,193,128 |
22,275,604 |
Group III: Acidic amino acids |
 |
| 4. Lysine |
313° |
255,128,227 |
55,283,264 |
Group IV: Basic amino acids |
 |
| 5. Cysteine |
63° |
249,255,128 |
33,877,310 |
Group II: Polar, uncharged amino acids |
 |
| 6. Glycine |
78° |
217,255,128 |
49,775,292 |
Group I: Nonpolar amino acids |
 |
| 7. Alanine |
94° |
183,255,128 |
40,519,668 |
Group I: Nonpolar amino acids |
 |
| 8. Valine |
125° |
128,255,138 |
49,100,261 |
Group I: Nonpolar amino acids |
 |
| 9. Leucine |
141° |
128,255,172 |
104,225,026 |
Group I: Nonpolar amino acids |
 |
| 10. Isoleucine |
157° |
128,255,206 |
58,789,793 |
Group I: Nonpolar amino acids |
 |
| 11. Phenylalanine |
172° |
128,255,238 |
56,025,389 |
Group I: Nonpolar amino acids |
 |
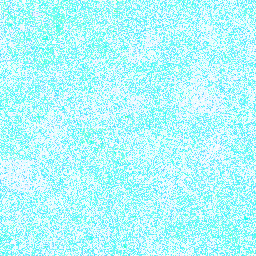
| 12. Tryptophan |
188° |
128,238,255 |
18,016,570 |
Group I: Nonpolar amino acids |
 |
| 13. Serine |
203° |
128,206,255 |
85,648,033 |
Group II: Polar, uncharged amino acids |
 |
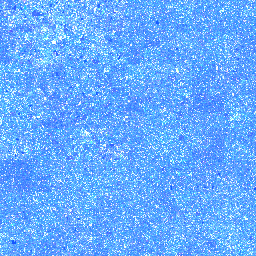
| 14. Threonine |
219° |
128,172,255 |
48,993,474 |
Group II: Polar, uncharged amino acids |
 |
| 15. Glutamine |
250° |
149,128,255 |
37,231,115 |
Group II: Polar, uncharged amino acids |
 |
| 16. Asparagine |
266° |
183,128,255 |
38,119,502 |
Group II: Polar, uncharged amino acids |
 |
| 17. Tyrosine |
282° |
217,128,255 |
32,762,873 |
Group II: Polar, uncharged amino acids |
 |
| 18. Arginine |
297° |
249,128,255 |
48,109,139 |
Group IV: Basic amino acids |
 |
| 19. Proline |
344° |
255,128,162 |
49,830,352 |
Group I: Nonpolar amino acids |
 |
| 20. Methionine |
110° |
149,255,128 |
18,447,715 |
START Codon |
 |
| 21. Ochre |
0° |
255,128,128 |
19,746,126 |
STOP Codon |
 |
| 22. Amber |
47° |
255,227,128 |
12,725,300 |
STOP Codon |
 |
| 23. Opal |
240° |
128,128,255 |
18,448,575 |
STOP Codon |
 |